  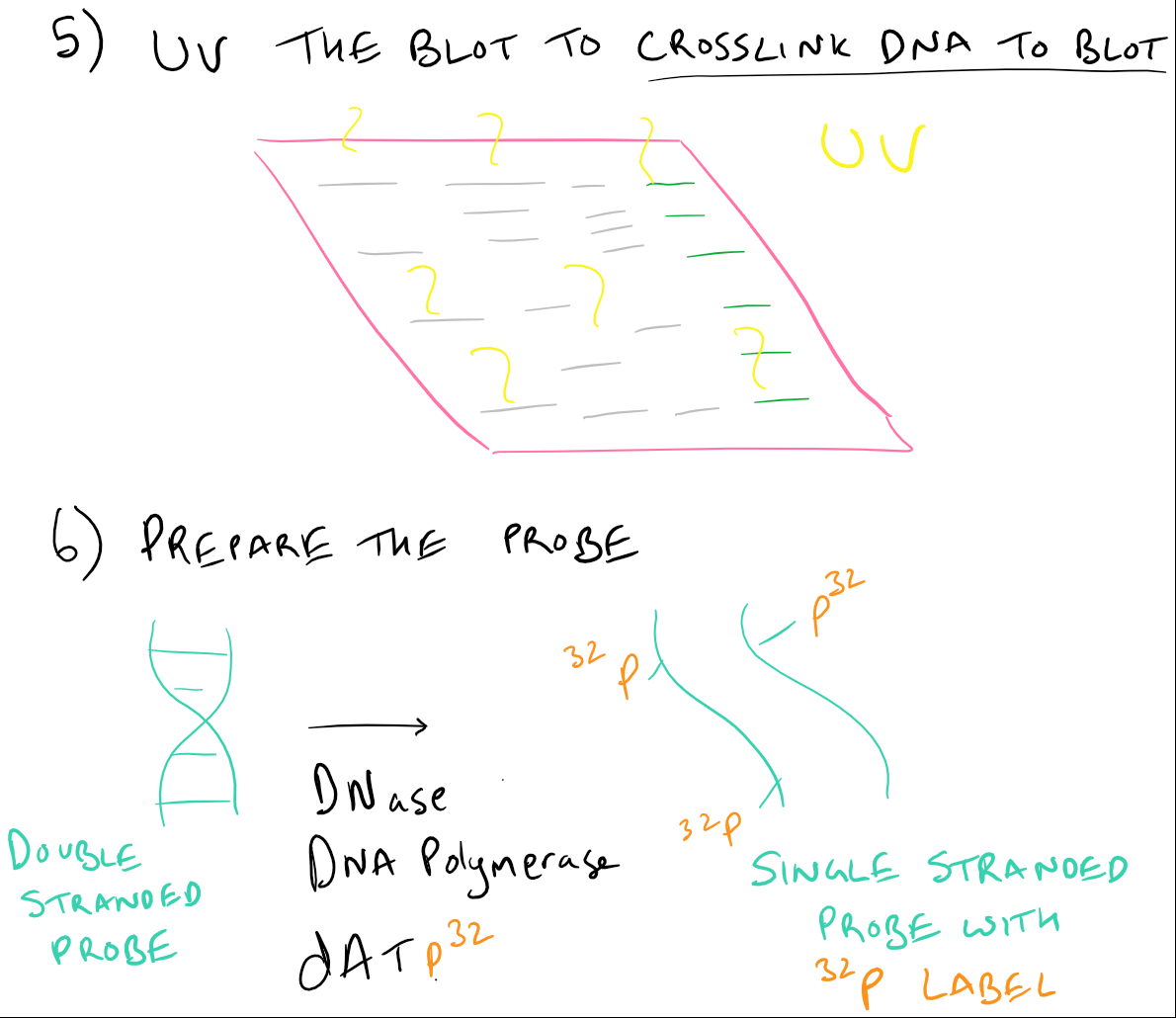
The DNA molecules which are separated get denatured by exposure to mild alkali, and then further transferred to nylon paper or nitrocellulose.As DNA is negatively charged, it migrates towards the positively charged electrode (anode).The mixture is subjected to electrophoresis. The digest or mixture of DNA is loaded into a well of polyacrylamide or agarose gel.The genomic DNA is digested with a restriction enzyme (one or more) this DNA was isolated from cells/tissues.The process is briefly described in the following points: This is the first nucleic acid blotting technique. Used in DNA fingerprinting and to identify specific gene sequences.ĭefinition of Southern blotting technique George Stark's group in 1979 at Stanford University.Ĭamera, LED, cooled CCD, film, or infrared imaging system. The most widely and accepted technique that identifies the sequence of amino acid in the protein mixture of the given sample. The technique that identifies a specific sequence of RNA or mRNA molecules for gene expression in the given blood sample. The technique used in the laboratory to detect the specific sequence of DNA in the provided tissue or blood sample. Content: Southern Vs Northern Vs Western Blotting Techniques In this article, we will observe the critical differences that lie between these three techniques, along with their description. The general process that is followed in blotting is in simple four steps, and firstly the sample is homogenized secondly, separation of target molecules is done by an electrophoresis membrane thirdly transferring of molecules to nylon or nitrocellulosic membrane and lastly, identification or hybridization of molecules is made. Therefore, these types of blotting like Southern, Northern and Western are the subtypes of blotting and rely on the target molecule that is being separated. It is a hybridization technique for the identification of specified genes, proteins or nucleic acid (DNA, mRNA) during multiple stages of gene expression. This technique involves immobilization of the sample of molecules of nucleic acid on the support of nylon membranes or nitrocellulose. Western blotting is a sensitive assay used in the detection of proteins in the given sample, and Stark’s group developed it.īlotting techniques are a widely used method for the specific identification of desired nucleic acid (DNA and RNA fragments) and proteins from the numerous molecules. Wood WI, Gitschier J, Lasky LA, Lawn RM (1985) Base composition-independent hybridization in tetramethylammonium chloride: a method for oligonucleotide screening of highly complex gene libraries.Northern blotting is the technique used to know the gene expression of the provided sample by identifying the specific RNA or mRNA molecules it was developed by David Kemp, James Alwine and George Stark in 1977 at Stanford University. Thomas PS (1980) Hybridization of denatured RNA and small DNA-fragments transferred to nitrocellulose. Southern EM (1975) Detection of specific sequences. A Laboratory Manual (2nd ed), Cold Spring Harbor. Sambrook J, Fritsch T, Maniatis (1989) Molecular cloning. In: Rolfs A, Schumacher HC, Marx P (eds) PCR Topics. Rüger R, Höltke H-J, Reischl R, Sagner G, Kessler C (1991) Labeling of specific DNA-sequences with digoxigenin during polymerase chain reaction. Non-radioactive detection of differentially expressed genes using complex RNA or DNA hybridization probes. Ross R, Ross X, Rüger B, Laengin T, Reske-Kunz A. Lahaye T, Rueger B, Toepsch S, Thalhammer J and Schulze-Lefert P (1996) Detection of single-copy sequences with Digoxigenin-labeled probes in a complex plant genome after separation on pulsed-field gels. This process is experimental and the keywords may be updated as the learning algorithm improves.Ĭhurch G, Gilbert W (1984) Genomic sequencing. These keywords were added by machine and not by the authors. How to prepare DNA or RNA agarose gels is described elsewhere (see Sambrook et al., 1989). For dot blot hybridization, DNA or RNA is spotted directly onto a membrane, while for Southern or northern blot hybridization DNA fragments or mRNAs, respectively, are transferred to the membrane after size separation on an agarose gel by capillary-, vacuum-, pressure-or electroblotting and subsequently hybridized with a labeled probe that can detect a specific sequence. 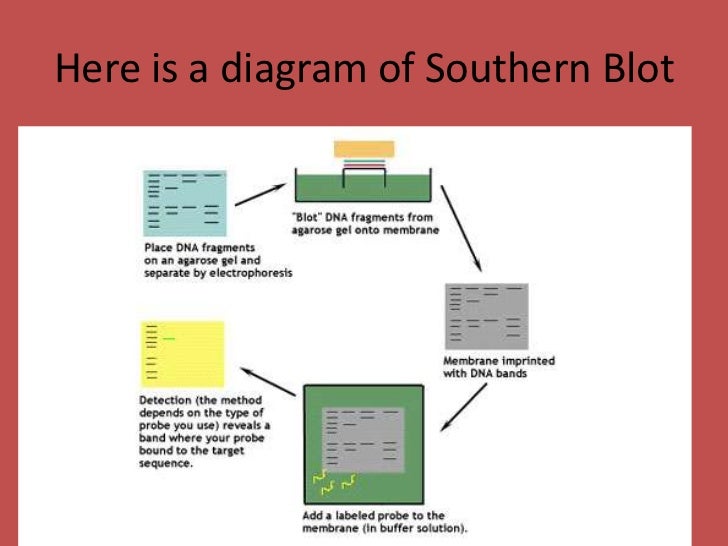
In principle molecular hybridization is the formation of double-stranded nucleic acid molecules by sequence-specific base pairing of complementary single strands. Here, we will specifically deal with applications of the digoxigenin (DIG) system in dot blot, Southern, and northern blot hybridizations. Nucleic acid hybridization on solid supports (nitrocellulose or nylon membranes) has become one of the most important techniques in molecular biology. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
